
Funding

“We use mathematics, computing and statistics to explore the networks of biochemical reactions inside our cells”
I am broadly interested in understanding what enables cells to grow, divide, transmit signals, process information and make decisions. These processes are particularly interesting because they happen reliably in spite of the inherent randomness, or stochasticity, of the biochemical networks inside the cell.
The goal of my research is to identify the principles that govern the design and robust function of networks of interacting genes and proteins. These design principles are ones that have been consistently chosen by evolution, and so may play an important role in the cell.
Biological networks are too complex to be understood by studying their individual components alone. To study the networks at a systems-level it is necessary to use a theoretical approach. To do this I use tools from physics, mathematics and bioinformatics. This allows me to explore the temporal, spatial and stochastic dynamics of biochemical network models using analytical, computational or statistical and computational methods.
Experimental advances in high-throughput methods in molecular biology provide systems-level data. Single cell studies quantify variations in biomolecular copy numbers in the cells and their corresponding effects on the cell’s appearance and characteristics, it’s phenotype. Together, these data provide an excellent opportunity for system-level and quantitative study of the mechanisms of robust cellular function, and phenotypic variability. To this end, I regularly collaborate with biologists that generate system level and single cell level experimental data to incorporate this into our mathematical models.
I am based in the Department of Mathematics at Imperial College London. I am also the mathematician in residence at MRC Laboratory of Medical Sciences (MRC LMS), where I consult and collaborate with MRC LMS researchers on the application of mathematical approaches in biomedical science.
Read more about Shahrezaei’s work on using maths to model the “bet hedging” of biology.
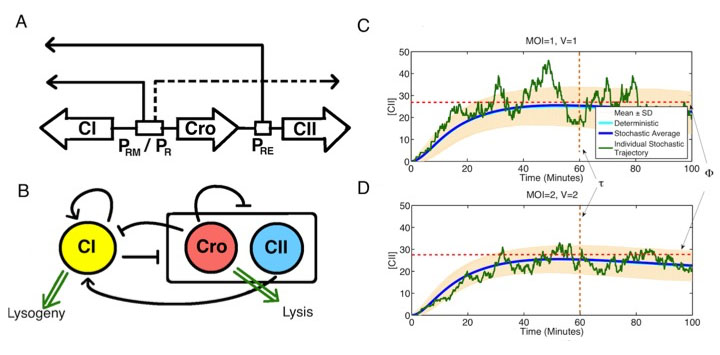
Stochastic decision making in the lambda phage (A-B) Schematic of the core genetic network involved in lysis-lysogeny decision. CI gene promotes itself and represses the other genes. Cro represses everything, while CII promotes CI. (C-D) It is observed that multiplicity of infection (MOI) can affect decision making even if the concentration of phage in bacteria is remained constant. Stochastic simulations of this genetic system shows that one phage in a small cell compared to two phage in cell with double volume produces very similar mean [CII] time course (blue lines). However, the stochastic trajectory of [CII] (green lines) is much more variable in the MOI= 1 case, resulting in a higher chance of crossing threshold (dashed lines) and inducing lysogeny fate.
Selected Publications
Bertaux F, Marguerat S, Shahrezaei V. (2018). Division rate, cell size and proteome allocation: impact on gene expression noise and implications for the dynamics of genetic circuits. Royal Society Open Science 5.
Uhia I, Priestman M, Joyce G, Krishnan N, Shahrezaei V, Robertson BD. (2018). Analysis of ParAB dynamics in mycobacteria shows active movement of ParB and differential inheritance of ParA. PLOS One 13.
Priestman M, Thomas P, Robertson BD, Shahrezaei V. (2017). Mycobacteria Modify Their Cell Size Control under Sub-Optimal Carbon Sources. Front Cell Dev Biol 5.
Sturrock M, Li S, Shahrezaei V. (2017). The influence of nuclear compartmentalisation on stochastic dynamics of self-repressing gene expression. Journal of Theoretical Biology 424, 55-72.
Archer N, Walsh MD, Shahrezaei V, Hebenstreit D. (2016). Modeling Enzyme Processivity Reveals that RNA-Seq Libraries Are Biased in Characteristic and Correctable Ways. Cell Systems 3, 467.
Shahrezaei V, Marguerat S. (2015). Connecting growth with gene expression: of noise and numbers. Curr Opin Microbiol 25, 127-135.
Robb M, Shahrezaei V. (2014). Stochastic cellular fate decision making by multiple infecting Lambda phage. PLOS One 9.
Dushek O, Lellouch A, Vaux D, Shahrezaei V. (2014). A novel biosensor architecture is predicted to reduce nonlinear reporting of signaling protein activity. Biophysical Journal 107, 773-82.
Szomolay B, Shahrezaei V. (2012). Bell-shaped and ultrasensitive dose-response in phosphorylation-dephosphorylation cycles: the role of kinase-phosphatase complex formation. BMC Systems Biology 6, 26.
Malleshaiah M*, Shahrezaei V*, Swain PS, Michnick SW. (2010). The Ste5 scaffold protein directly controls a switch-like mating decision in yeast. Nature 465, 101-105.
*Contributed equally
