The functional gene control group headed by Mikhail Spivakov investigates the logic of gene regulation using a combination of experimental and computational approaches.
“How molecular switch regions on the DNA work together to control gene expression and cell function?”
All cells in the body have the same DNA, but they look and function differently. This is because different parts of DNA are at work in different cells. These parts include not just genes but, importantly, also the ‘molecular switches’ on the DNA known as enhancers, which determine when genes should go on and off. Mutations in these regions are known to be behind both rare and common diseases, including developmental and autoimmune conditions and cancer. One of the puzzling things about enhancers is that they are often located on the DNA strands far away from the genes they control. However, the two meters of DNA are tightly packaged in the cells’ nuclei, which allows distal regions such as enhancers and genes to come into each other’s proximity. We aim to decipher the rules of gene control by enhancers and work out how they underscore some of the biggest decisions made by the cells. These decisions include, for example, whether a cell should divide or remain resting, what kind of cell a stem cell should develop into, and whether an immune cell should treat another cell as a friend or foe. In particular, we are interested in how contacts between enhancers and their target genes are made, why enhancers can be selective for some genes over others in the vicinity, and how enhancers jointly mediate concerted changes in the expression of multiple genes in response to environmental cues. Our work spans multiple spatial scales, with some research focused on the mechanisms linking enhancers with genes, through to studying large networks of enhancers and genes they control in specific biological contexts. We focus on the analyses of 3D DNA conformation, enhancer activity and gene expression in development, immune response and cancer, taking advantage of molecular methods that enable switching enhancers ‘on’ and ‘off’ and single-cell technologies. We employ advanced computational techniques to integrate and learn from the data generated using these experimental methods. A better understanding of how enhancers work in health and disease will enable us to better predict the effects of enhancer abnormalities on cellular and organismal function, as well as develop novel therapies that directly or indirectly target enhancers.
“Our group aims to understand how DNA regulatory elements such as gene enhancers regulate cellular transcriptional programmes, and how their function is modified by natural genetic variation. This is important because genetic variation at enhancers is known to underlie both rare and common diseases.”
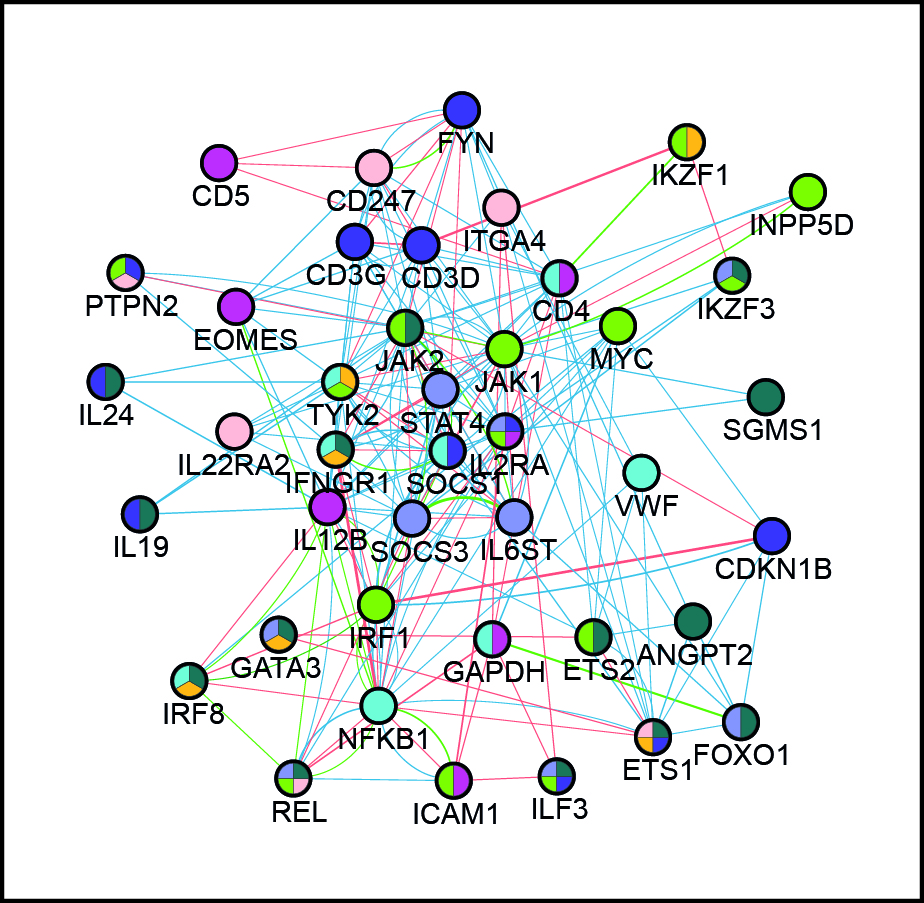

We aim to decipher the ground rules of gene control and establish their functional interplay in biological phenomena involving global changes in phenotype, such as in cell differentiation and activation. Our particular interest is in human primary cells as models, and we use genetic variation at enhancers as both experimental tools (“natural perturbations”) and objects of study in these systems. Our analyses span multiple scales, from pairwise enhancer-promoter interactions to cis-regulatory networks, from single cells to cell populations, and from single individuals to population cohorts. We combine experimental and computational approaches in our work, capitalising on our previous studies of promoter-enhancer relationships, organisation of DNA regulatory elements and population genomics. Our ultimate goal is to generate comprehensive functional models of gene control “logic” underlying cellular decisions. Interrogation and validation of these models will pinpoint key individual players (regulatory elements, genes, extrinsic signals) and their regulatory relationships in these processes and shed light on how they are remodelled in disease.
Our lab uses wet-lab and computational approaches to study how DNA regulatory elements, such as gene enhancers, control gene expression and how this goes wrong in disease, such as common pathologies of the immune and cardiovascular system and cancer.
As part of this research, we develop approaches to interpret the function of non-coding variants detected from GWAS studies using insights from statistical genetics, epigenomics and gene regulatory network analysis.
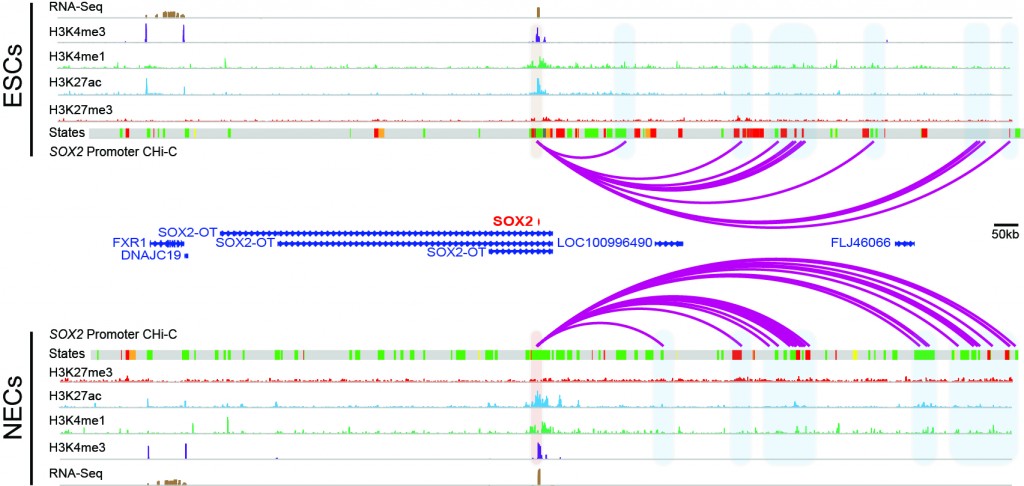
We are also interested in how enhancers find their target genes (which may be megabase pairs away from them), as well as the role these elements play in early development, cytokine response (with implications for autoimmunity) and cancer drug resistance.


Ray-Jones H^, Sung CK, Chan LT, Haglund A, Artemov P, Della Rosa M, Ruje L, Burden F, Kreuzhuber R, Litovskikh A, Weyenburgh E, Brusselaers Z, Tan VXH, Frontini M, Wallace C, Malysheva V*, Bottolo L*^, Vigorito E* & Spivakov M*^. Genetic coupling of enhancer activity and connectivity in gene expression control. Nat Commun,16, 970 (2025). https://doi.org/10.1038/s41467-025-55900-3 * joint supervisors, ^ joint corresponding authors.
Lambert J / Oc S, Worssam MD, Häußler D, Figg NL, Baxter R, Foote K, Finigan A, Mahbubani, KT, Bennett MR, Krüger A, Spivakov M, Jørgensen HF. Network-based prioritisation and validation of novel regulators of vascular smooth muscle cell proliferation in disease. Nature Cardiovascular Research, 3, 714–733, 2024
Ray-Jones H, Spivakov M. Transcriptional enhancers and their communication with gene promoters. (2021). Cell Mol Life Sci, doi: 10.1007/s00018-021-03903-w. (A review article).
Freire-Pritchett P* / Ray-Jones H*, Della Rosa M, Eijsbouts C, Orchard WR, Wingett SW, Wallace C, Cairns J, Spivakov M^, Malysheva V^. Detecting chromosomal interactions in Capture Hi-C data with CHiCAGO and companion tools. (2021). Nature Protocols, doi: 10.1038/s41596-021-00567-5. * joint first authors, ^ joint corresponding authors with a postdoc in the lab.
Watt S, Vasquez L, Walter K, Mann AL, Kundu K, Chen L, Yan Y, Ecker S, Burden F, Farrow S, Farr B, Iotchkova V, Elding H, Mead D, Tardaguila M, Ponstingl H, Richardson D, Datta A, Flicek P, Clarke L, Downes K, Pastinen T, Fraser P, Frontini M, Javierre BM^, Spivakov M^, Soranzo N^. (2021). Genetic perturbation in PU.1 binding and chromatin looping at neutrophil enhancers associates with autoimmune disease. Nature Communications 12:2298. ^ joint corresponding authors.
Ho JSY* / Mok BW-Y*, …, Ray-Jones H, Malysheva V, Thiecke MJ, …, Spivakov M, Weirauch MT, Heinz S, Chen H, Benner C, Richt JA, Marazzi I. (2021). TOP1 inhibition therapy protects against SARS-CoV-2-induced lethal inflammation. Cell 184, 1–15. * joint first authors.
Thiecke MJ*/ Wutz G*, Muhar M, Tang W, Bevan S, Malysheva V, Stocsits R, Neumann T, Zuber J, Fraser P, Schoenfelder S, Peters J-M, Spivakov M. (2020). Cohesin-Dependent and -Independent Mechanisms Mediate Chromosomal Contacts between Promoters and Enhancers. Cell Reports 32: 107929. * joint first authors.
Dobnikar L* / Taylor AL*, Chappell J, Oldach P, Harman JL, Oerton E, Dzierzak E, Bennett MR, Spivakov M^ / Jørgensen HF^. (2018). Disease-relevant transcriptional signatures identified in individual smooth muscle cells from healthy mouse vessels. Nature Communications 9:4567. * joint first authors, ^ joint corresponding authors.
Freire-Pritchett P* / Schoenfelder S*, Várnai C, Wingett SW, Cairns J, Collier AJ, García-Vílchez R, Furlan-Magaril M, Osborne CS, Fraser P, Rugg-Gunn PJ^, Spivakov M^. (2017). Global reorganisation of cis-regulatory units upon lineage commitment of human embryonic stem cells. eLife 6: e21926. * joint first authors, ^ joint corresponding authors.
Javierre B-M* / Burren OS* / Wilder SP* / Kreuzhuber R* / Hill SM*, Sewitz S, Cairns J, Wingett SW, Várnai C, Thiecke MJ, Burden F, Farrow S, Cutler AJ, Rehnström K, Downes K, Grassi L, Kostadima M, Freire-Pritchett P, Wang F, The BLUEPRINT Consortium , Stunnenberg HG, Todd JA, Zerbino DR, Stegle O, Ouwehand WH, Frontini M^ / Wallace C^ / Spivakov M^# / Fraser P^. (2016). Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters Cell 167(5), 1369-1384. * joint first authors, ^ joint corresponding authors, # lead contact. • Minireview presenting this paper • Interview on Cambridge TV
Cairns J* / Freire-Pritchett P*, Wingett SW, Várnai C, Dimond A, Plagnol V, Zerbino D, Schoenfelder S, Javierre B-M, Osborne C, Fraser P, Spivakov M. (2016). CHiCAGO: Robust Detection of DNA Looping Interactions in Capture Hi-C data. Genome Biology 17(1), 127. PMID: 27306882. F1000 Prime recommended paper. * joint first authors.
Bolland DJ* / Koohy H*, Wood AL, Matheson LS, Krueger F, Stubbington MJT, Baizan-Edge A, Chovanec P, Stubbs BA, Tabbada K, Andrews SR, Spivakov M^, Corcoran AE^. (2016). Two Mutually Exclusive Local Chromatin States Drive Efficient V(D)J Recombination. Cell Reports 15(11), 2475–2487. PMID: 27264181. F1000 Prime recommended paper. * joint first authors; ^ joint corresponding authors.
Spivakov M. (2014). Spurious transcription factor binding: Non-functional or genetically redundant? BioEssays 36(8), 798-806. PMID: 24888900.
Junion G* / Spivakov M*, Girardot C, Braun M, Gustafson EH, Birney E, Furlong EE. (2012). A transcription factor collective defines cardiac cell fate and reflects lineage history. Cell 148(3):473-86. PMID: 22304916. * joint first authors.