By Helen Figueira
May 28, 2013
Time to read: 4 minutes
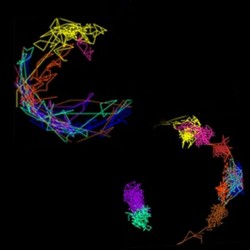 How chromosome movements dictate homolog pairing in meiosis
How chromosome movements dictate homolog pairing in meiosis
In the hunt for your soulmate, you can’t just settle for the first nice person you meet – you have to keep searching until you find that special someone who matches you perfectly. Chromosomes have to be similarly picky. Early in meiosis, the unique form of cell division that produces sperm and egg cells, chromosomes must find their exact pair and stick by them. New research from the CSC reveals that, like a singleton darting frantically around a speed dating session until they find a satisfactory suitor, the movement of these chromosomes during their search is crucial to their success in finding their partner.
At the start of meiosis, cells have two versions of each chromosome – one inherited from father, one from mother. For meiosis to advance, they must each locate and join to their complementary, or homologous, partner. Enrique Martinez-Perez (Meiosis Group), who led the study published in PLoS Genetics, wants to know how this happens. “There is a lot of chromosome movement at the stage when homologous pairing occurs,” he says. “But there’s been much debate over whether this movement was promoting homolog encounters, or preventing incorrect match-ups.”
“When pairs come together, a structure called the synaptonemal complex (SC) forms. This is like a glue to hold homologs together,” explains Martinez-Perez. Previous studies had investigated the effects of movement by tinkering with the proteins that drive the action. Proteins straddling the outer casing of the nucleus join the chromosomes inside to motor proteins outside, which run along the cell’s cytoskeleton. As the motor proteins move, they drag the chromosomes around the nucleus like a puppet on a string.
“Instead of looking at organisms with mutations in these proteins, which has been done already , we used a mitochondrial mutant,” he reveals. This effectively removed the motors’ fuel, leaving them stationary. “We could then dissect the effect of movement specifically, without altering the structure of the proteins.”
The result was that the wrong chromosomes glued together. Previously it had been supposed that in meiosis homologs drifted together, and the motor-driven movement ripped them apart; only homologous pairs would be able to resist this, and only then would SC formation be licensed. “However, this is not the case,” argues Martinez-Perez, “because in our mutants there is almost no motor-driven movement, but the SC can still be built.”
Motor driven movement is required to make sure each chromosome can encounter its homolog. It then has a role in quality control. The prevalence of mismatched pairing in the mutants implies that the movement is in some way required to ensure that only the right pairs form a bond. “How homology is assessed and SC formation is triggered remains largely unknown,” admits Martinez-Perez, “but we now know that movement is essential for the mechanism that promotes homology search.’
This fundamental research was based on the model organism C. elegans, but the practical importance is clear: “We know these processes are conserved across species, so have a direct relevance to human biology. Pairing problems leads to aneuploid gametes, which in humans can cause spontaneous abortions or Down’s syndrome. Understanding how homologs pair at the start of meiosis is a key question in the field, but despite the importance, it remains poorly understood.”
AL
Reference: Labrador, L., Barroso, C., Lightfoot, J., Müller-Reichert, T., Flibotte, S., Taylor, J., Moerman, D. G., Villeneuve, A. M., Martinez-Perez, E., (2013). Chromosome movements promoted by the mitochondrial protein SPD-3 are required for homology search during caenorhabditis elegans meiosis. PLoS genetics 9 (5).
Chromosome movement around the nuclear envelope during meiosis in healthy wild-type cells (left) and mutants.