CRISPR is a powerful DNA-editing tool that has underpinned huge advancements in human healthcare in the last decade. It is a precision tool, but is not perfect, and misplaced DNA edits can compromise safety and efficacy, costing billions each year. Researchers at the MRC Laboratory of Medical Sciences (LMS), Imperial College London and the University of Sheffield have published research in Nature showing that the physical twisting of DNA plays an important role in these mistakes. Using a newly developed platform of tiny (nanometer-sized) DNA circles, called DNA minicircles, the team captured never-before-seen interactions between CRISPR and DNA, providing insights that could help eradicate errors altogether.
By Emily Armstrong
March 25, 2026
Time to read: 8 minutes
CRISPR-Cas9 has transformed biology by giving scientists a programmable way to cut and edit DNA. Its ever-growing impact includes groundbreaking therapies for genetic diseases such as sickle cell anaemia and an increasing role in personalised cancer treatment and rapid diagnostics. But even carefully designed CRISPR systems can sometimes cut DNA sequences that were not the intended targets. “It’s a tool that is not perfect and can introduce errors and make edits where it shouldn’t make them,” says Professor David Rueda, head of the Single Molecule Imaging group at the LMS and Chair in Molecular and Cellular Biophysics at Imperial College London. “And it’s an important problem for the industry. It’s been estimated to be $0.3 to $0.9 billions per year in industry costs, in profiling off-targets, redesigning guides and delays.”
Most research into CRISPR accuracy has focused on the DNA sequence the system recognises, but recent work has shown that the shape, or topology, of DNA molecules can be as important as the sequence itself. In the body, DNA is constantly being bent, stretched and twisted as genes are transcribed and replicated. One common state is negative supercoiling, where the DNA helix becomes slightly underwound. Because the molecule’s ends cannot freely rotate, the DNA can relieve this stress by buckling into loops—much like a spring that coils up when it is slightly untwisted.
“Several labs, including ours and Sheffield’s, have shown that DNA topology is important,” says David. “But the question was: how does Cas9 interact with DNA in the supercoiled state?” Answering that question required an entirely new experimental approach.
Quentin Smith spent the first year of his PhD designing a new experimental system: DNA minicircles that can be deliberately supercoiled. These tiny loops of DNA mimic the torsional stress that DNA experiences inside cells. But creating them at the right scale was not straightforward.
“Minicircles existed before, but they were too large to produce a clear structure in cryo-electron microscopy,” says Quentin. “The challenge was to make them as small as possible to reveal the full Cas9-DNA structure, while still allowing them to be supercoiled. If they’re too small, that becomes energetically impossible. It’s a bit like a Goldilocks problem – not too small, not too big, but just right.”
By finding this sweet spot, Quentin created DNA circles that were small enough to visualise by cryo-electron microscopy and that maintained a supercoiled state.
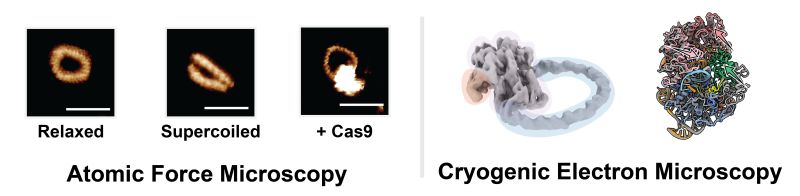
They collaborated with the team at Sheffield to use their high-resolution atomic force microscopy technique to determine the sweet spot where the DNA circles were small enough to visualise and still maintained a supercoiled state. Professor Alice Pyne, Head of the Royce Nanocharacterisation Laboratory at the University of Sheffield, said, “We were able to image Quentin’s DNA minicircles in solution, observing their helical structure as they buckled under stress in their supercoiled state. These minicircles are smaller than anything we’ve been able to previously create, pushing the limits of our microscopy technologies.”
Collaborators at Sheffield were able to visualise these supercoiled DNA minicircles down to their helical structure using high-resolution atomic force microscopy, showing that there were differences in how Cas9 behaved depending on the DNA sequence and the topology. Sylvia Whittle, the lead Sheffield author developed new machine learning tools to quantify the changes they observed in their data. “By visualising and quantifying changes in the DNA helical structure, we were able to gain fresh insight into how effective different Cas9 interactions can be.”
The team found that supercoiling strands of DNA makes them far more susceptible to unintentional cutting: “The same DNA with the same sequence, in linear form, is not cut by Cas9. It remains completely intact,” says David. “But if you supercoil it, now it’s cut by Cas9. We think this means that many of the off-targets observed in cells appear not only because of the sequence, but also because the DNA is supercoiled.”
“Using these minicircles, Quentin was able to obtain a high-resolution structure of Cas9 on supercoiled DNA. You can see the entire Cas9 and how it fits with the DNA. There are very few examples of enzymes bound to supercoiled complexes in the literature.”
This allowed the researchers to see, at near-atomic resolution, how Cas9 interacts with supercoiled DNA. The images revealed something striking: when Cas9 binds to supercoiled DNA, the enzyme changes its geometry to prime itself to cut the DNA strand. It moves the part of the enzyme responsible for cutting the target DNA strand – the HNH domain – closer to the point where CRISPR cuts the DNA, readying the enzyme to cut.
Their hypothesis is that as DNA twists and buckles, it reduces the energy required to unzip, lowering the energy barrier for Cas9 binding and cutting, and thereby facilitating off-target activity.
When the team compared the molecular structures of Cas9 that they had developed with those previously published, the differences were dramatic. Most earlier structures were obtained based on how Cas9 interacts with linear DNA, so those models lack the realistic mechanical stresses found in coiled DNA in living cells. The new versions of the structure proposed by this research reveal that supercoiling fundamentally changes how the enzyme behaves.
At the same time, the researchers saw for the first time that the twisting of the DNA helix during supercoiling makes mismatches easier for Cas9 to tolerate. Imperfect sequence matches between the CRISPR guide and DNA would normally stop Cas9 from working, but the images of Cas9 bound to supercoiled DNA revealed new types of mismatches being made, showing new mechanisms of off-target cutting.
“We knew topology played a role in mismatches,” says David. “But how the protein and DNA interact at the molecular level to cause this was not known. These structures now reveal that.”
Understanding the biomechanics of how supercoiling affects CRISPR accuracy could have major implications for gene-editing technologies.
Current strategies for developing high-fidelity, low-error CRISPR precision rely heavily on structural information obtained from linear DNA experiments. But if Cas9 behaves differently on supercoiled DNA, those designs may not fully capture what happens inside living cells. Quentin believes the new findings open the door to developing CRISPR enzymes that can account for DNA topology and dramatically reduce errors in the system. Achieving that step would have a huge impact on healthcare development.
He says, “This study definitely paves the way to generate Cas9 variants that are sensitive to topology. Most high-fidelity variants were designed using linear DNA structures. But in cells, the DNA is supercoiled to different degrees, so you might not get the same reduction in off-target activity in the body that you see in the lab.”
David says, “It’s amazing, we all take Cas9 for granted and think we know everything about it. But we still haven’t seen the truly active structure. This work takes us one step closer – and it paves the way for developing new, more accurate variants.”
This project is a testament to the importance of bringing together diverse perspectives and backgrounds. David says, “My lab is not a structural biology lab. We had never done structural biology before. Quentin’s drive and enthusiasm were essential in developing the programme in my lab, with the support of others at the LMS, including our Cryo-Electron Microscopy Facility Manager Dr Ricardo Aramayo, Dr Adam Jalal, Ms Deborah Egharevba, Dr Kyle Morris (EMBL-EBI) and Dr Nora Cronin at Imperial’s London Consortium for Cryo-Electron Microscopy (LonCEM).”
With new tools to study DNA under realistic physical conditions, the researchers believe that many more discoveries lie ahead – not only for Cas9, but for a wide range of DNA-binding proteins. As Quentin says, “Now that we can use these supercoiled minicircles, we can start uncovering many different things, not just for Cas9, but potentially for other CRISPR systems and DNA-binding proteins as well.”
This study was funded by the Medical Research Council, the Engineering and Physical Sciences Research Council and a UKRI Future Leaders Fellowship.
Quentin received a Student Equipment Access Scheme, enabling access to Bruker Dimension XR facilities at The Royce Discovery Centre at the University of Sheffield.
This article was written by Anthony Lewis.
Read the full publication: https://www.nature.com/articles/s41586-026-10255-7