“How is DNA controlled and regulated?”
DNA is not a static molecule – proteins and other molecules constantly interact with its strands to unwind them and express its genes. This dynamic network of compounds surrounding DNA is called the epigenome, and the compounds are constantly changing and moving to ensure genes are expressed when and where they are needed.
Some of these molecules are transcription factors; proteins that attach to DNA to turn genes “on” and “off.” They attach to portions of DNA called gene promoters, which reside upstream to genes and allow the factors to begin transcription.
Transcription regulatory networks (TRNs) are how these transcription factors work together with the genes they regulate.
Our group is studying this epigenomic activity is regulated, with a focus on specific elements including gene promoters, transcription factors, and TRNs.
Our group studies gene regulation at a genome-wide level using computational modelling, with our most recent work focussed on how we classify core promoters, how we can better match regulatory elements to the genes they influence, and how the epigenome guides development.
Most of our work is computational – we analyse previously published data or work in partnership with experimental laboratories. We also carry out our own experimental work focussing on how promoters affect development and differentiation.
“Our research investigates how regulatory information is encoded in DNA, how it is interpreted and how it is communicated across cells in multicellular organisms”
The development of complex organisms critically depends upon regulation of gene activity across dozens, hundreds or millions of cells in time and space. This regulation ensures the fidelity of development of complex structures and the overall function of the organism. Its disruption therefore often leads to disease.
We study gene regulation at the genome-wide level using computational genomics and epigenomics, as well as experimental transcriptomics and promoter activity mapping. Our major research interests focus on:
Our recent work centres on the functional classification of core promoters, the development of methods to correctly assign regulatory elements to the genes they regulate, and the role of the epigenome in guiding development. Most of our work until now has been purely computational, either analysing publicly available data or in collaboration with experimental laboratories using high-throughput sequencing techniques such as CAGE, ATAC-seq, MNase-seq, RNA-seq, ChIP-seq and Hi-C. Since 2016 we also have experimental activity focusing on studying functional diversity of promoters in development and differentiation.
You can find out more on their independent Computational Regulatory Genomics website
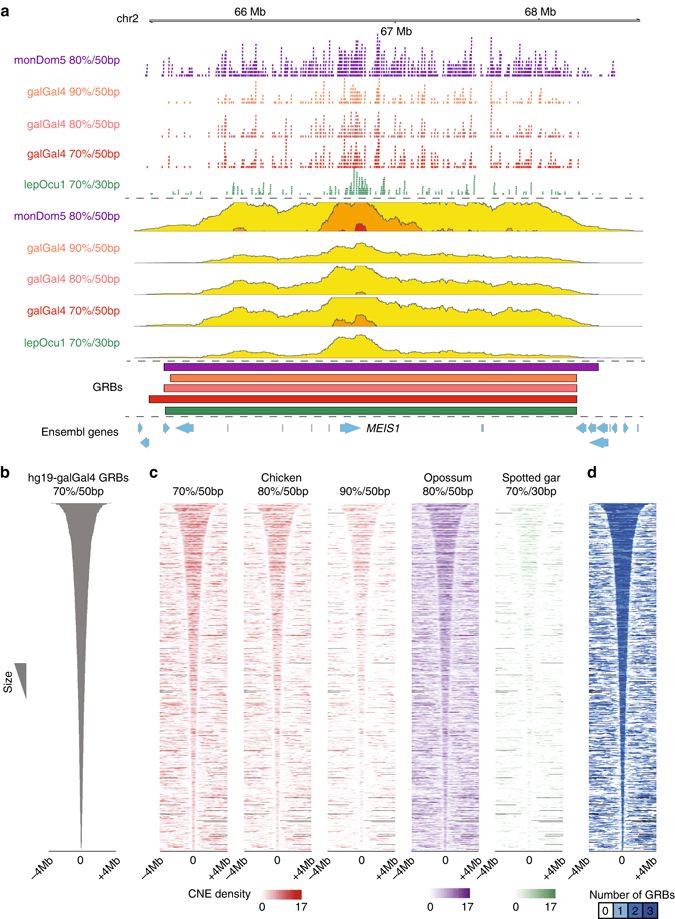
We investigate how genes are regulated during development, differentiation and disease, using computational and experimental genomics. We study how promoters and enhancers cooperate to control transcription, and how the architecture of regulatory landscapes enables precise patterns of gene expression across tissues and developmental stages. A particular focus is on long-range regulation, where clusters of enhancers spanning hundreds of kilobases interact with a single developmental gene within structures known as genomic regulatory blocks (GRBs).
This research has direct relevance to human disease. Many genetic variants associated with complex disorders lie in non-coding regions, often far from the genes they affect. By modelling the organisation of enhancers within GRBs, we have shown that disease-associated mutations in type 2 diabetes, obesity, and schizophrenia act on distant developmental regulators rather than on the nearest genes. This has led to the identification of previously unrecognised disease mechanisms involving long-range misregulation of key transcription factors.
Going forward, we aim to develop innovative computational approaches of analysing multiomic and comparative data to understand how disruptions of long-range regulatory architecture contribute to heritable developmental disorders, particularly those affecting the nervous system and body patterning and applying it to clinical data.





Wragg JW, Roos L, Vucenovic D, Cvetesic N, Lenhard B and Müller F. (2020). Embryonic tissue differentiation is characterized by transitions in cell cycle dynamic-associated core promoter regulation. Nucleic Acids Research, https://doi.org/10.1093/nar/gkaa563
Lenhard B and Sternberg MJE. (2020). Computational Resources for Molecular Biology: Special Issue 2020. Journal of Molecular Biology, 431(13), 2395.
Bonetti A, Agostini F, Suzuki AM, Hashimoto K, Pascarella G, Gimenez J, Roos L, Nash AJ, Ghilotti M, Cameron CJF, Valentine M, Medvedeva YA, Nogucki S, Agirre E, Kashi K, Luginbuhl J, Cazzoli R, Agrawal S, Luscombe NM, Blanchette M, Kasukawa T, De Hoon M, Arner E, Lenhard B, Plessy C, Castelo-Branco G, Orlando V, Carninci P. (2020). RADICL-seq identifies general and cell type–specific principles of genome-wide RNA-chromatin interactions. Nature Communications, doi.org/10.1038/s41467-020-14337-6.
Nepal C, Hadzhiev Y, Balwierz P, Tarifeno-Saldivia E, Cardenas R, Wragg JW, Suzuki A-M, Carninci P, Peers B, Lenhard B, Andersen JB, Muller F. (2020). Dual-initiation promoters with intertwined canonical and TCT/TOP transcription start sites diversify transcript processing. Nature Communications, 11(168).
Fornes O, Castro-Mondragon JA, Khan A, van der Lee R, Zhang X, Richmond PA, Modi BP, Correard S, Gheorghe M, Baranasic D, Santana-Garcia W, Tan G, Cheneby J, Ballester B, Parcy F, Sandelin A, Lenhard B, Wasserman WW, Mathelier A. (2020). JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acid Research, doi.org/10.1093/nar/gkz1001.
Baresic A, Nash AJ, Dahoun T, Howes O and Lenhard B. (2020). Understanding the genetics of neuropsychiatric disorders: the potential role of genomic regulatory blocks. Nature Molecular Psychiatry 25, 6–18.
Nash AJ and Lenhard B. (2019). A novel measure of non-coding genome conservation identifies genomic regulatory blocks within primates. Bioinformatics 35 (14), 2354–2361.
Newton MD, Taylor BJ, Driessen RPC, Roos L, Cvetesic N, Allyjaun S, Lenhard B, Cuomo ME & Rueda DS. (2019). DNA stretching induces Cas9 off-target activity. Nature Structural & Molecular Biology, doi: 10.1038/s41594-019-0188-z
Börlin CS, Cvetesic N, Holland P, Bergenholm D, Siewers V, Lenhard B, Nielsen J. (2018). Saccharomyces cerevisiae displays a stable transcription start site landscape in multiple conditions. FEMS Yeast Research, doi.org/10.1093/femsyr/foy128
Nash AJ and Lenhard B. (2018). A novel measure of non-coding genome conservation identifies genomic regulatory blocks within primates. Bioinformatics, doi: 10.1093/bioinformatics/bty1014
Cvetesic N, Leitch HG, Borkowska M, Muller F, Carninci P, Hajkova P, Lenhard B. (2018). SLIC-CAGE: high-resolution transcription start site mapping using nanogram-levels of total RNA. Genome Research, doi:10.1101/gr.235937.118
Khan A, Fornes O, Stigliani A, Gheorghe M, Castro-Mondragon JA, van der Lee R, Bessy A, Chèneby J, Kulkarni SR, Tan G, Baranasic D, Arenillas DJ, Sandelin A, Vandepoele K, Lenhard B, Ballester B, Wasserman WW, Parcy F, Mathelier A. (2017). JASPAR 2018: update of the open-access database of transcription factor binding profiles and its web framework. Nucleic acids research 46(D1), D260-D266.
Harmston N, Ing-Simmons E, Tan G, Perry M, Merkenschlager M, Lenhard B. (2017). Topologically associating domains are ancient features that coincide with Metazoan clusters of extreme noncoding conservation. Nature communications 8(1), 441.
Polychronopoulos D, King JWD, Nash AJ, Tan G, Lenhard B. (2017). Conserved non-coding elements: developmental gene regulation meets genome organization. Nucleic acids research 45(22), 12611-12624.
Cvetesic N, Lenhard B. (2017). Core promoters across the genome. Nature biotechnology 35(2), 123.
Haberle V, Forrest AR, Hayashizaki Y, Carninci P, Lenhard B. (2015). CAGEr: precise TSS data retrieval and high-resolution promoterome mining for integrative analyses. Nucleic Acids Res pii: gkv054. [Epub ahead of print].
Ing-Simmons E, Seitan VC, Faure AJ, Flicek P, Carroll T, Dekker J, Fisher AG, Lenhard B, Merkenschlager M. (2015). Spatial enhancer clustering and regulation of enhancer-proximal genes by cohesin. Genome Res [Epub ahead of print].
FANTOM Consortium and the RIKEN PMI and CLST (DGT), Forrest AR, Kawaji H, Rehli M, Baillie JK, de Hoon MJ, Haberle V, Lassmann T, Kulakovskiy IV, Lizio M, Itoh M, Andersson R, Mungall CJ, Meehan TF, Schmeier S, Bertin N, Jørgensen M, Dimont E, Arner E, Schmidl C, Schaefer U, Medvedeva YA, Plessy C, Vitezic M, Severin J, Semple C, Ishizu Y, Young RS, Francescatto M, Alam I, Albanese D, Altschuler GM, Arakawa T, Archer JA, Arner P, Babina M, Rennie S, Balwierz PJ, Beckhouse AG, Pradhan-Bhatt S, Blake JA, Blumenthal A, Bodega B, Bonetti A, Briggs J, Brombacher F, Burroughs AM, Califano A, Cannistraci CV, Carbajo D, Chen Y, Chierici M, Ciani Y, Clevers HC, Dalla E, Davis CA, Detmar M, Diehl AD, Dohi T, Drabløs F, Edge AS, Edinger M, Ekwall K, Endoh M, Enomoto H, Fagiolini M, Fairbairn L, Fang H, Farach-Carson MC, Faulkner GJ, Favorov AV, Fisher ME, Frith MC, Fujita R, Fukuda S, Furlanello C, Furino M, Furusawa J, Geijtenbeek TB, Gibson AP, Gingeras T, Goldowitz D, Gough J, Guhl S, Guler R, Gustincich S, Ha TJ, Hamaguchi M, Hara M, Harbers M, Harshbarger J, Hasegawa A, Hasegawa Y, Hashimoto T, Herlyn M, Hitchens KJ, Ho Sui SJ, Hofmann OM, Hoof I, Hori F, Huminiecki L, Iida K, Ikawa T, Jankovic BR, Jia H, Joshi A, Jurman G, Kaczkowski B, Kai C, Kaida K, Kaiho A, Kajiyama K, Kanamori-Katayama M, Kasianov AS, Kasukawa T, Katayama S, Kato S, Kawaguchi S, Kawamoto H, Kawamura YI, Kawashima T, Kempfle JS, Kenna TJ, Kere J, Khachigian LM, Kitamura T, Klinken SP, Knox AJ, Kojima M, Kojima S, Kondo N, Koseki H, Koyasu S, Krampitz S, Kubosaki A, Kwon AT, Laros JF, Lee W, Lennartsson A, Li K, Lilje B, Lipovich L, Mackay-Sim A, Manabe R, Mar JC, Marchand B, Mathelier A, Mejhert N, Meynert A, Mizuno Y, de Lima Morais DA, Morikawa H, Morimoto M, Moro K, Motakis E, Motohashi H, Mummery CL, Murata M, Nagao-Sato S, Nakachi Y, Nakahara F, Nakamura T, Nakamura Y, Nakazato K, van Nimwegen E, Ninomiya N, Nishiyori H, Noma S, Noma S, Noazaki T, Ogishima S, Ohkura N, Ohimiya H, Ohno H, Ohshima M, Okada-Hatakeyama M, Okazaki Y, Orlando V, Ovchinnikov DA, Pain A, Passier R, Patrikakis M, Persson H, Piazza S, Prendergast JG, Rackham OJ, Ramilowski JA, Rashid M, Ravasi T, Rizzu P, Roncador M, Roy S, Rye MB, Saijyo E, Sajantila A, Saka A, Sakaguchi S, Sakai M, Sato H, Savvi S, Saxena A, Schneider C, Schultes EA, Schulze-Tanzil GG, Schwegmann A, Sengstag T, Sheng G, Shimoji H, Shimoni Y, Shin JW, Simon C, Sugiyama D, Sugiyama T, Suzuki M, Suzuki N, Swoboda RK, ‘t Hoen PA, Tagami M, Takahashi N, Takai J, Tanaka H, Tatsukawa H, Tatum Z, Thompson M, Toyodo H, Toyoda T, Valen E, van de Wetering M, van den Berg LM, Verado R, Vijayan D, Vorontsov IE, Wasserman WW, Watanabe S, Wells CA, Winteringham LN, Wolvetang E, Wood EJ, Yamaguchi Y, Yamamoto M, Yoneda M, Yonekura Y, Yoshida S, Zabierowski SE, Zhang PG, Zhao X, Zucchelli S, Summers KM, Suzuki H, Daub CO, Kawai J, Heutink P, Hide W, Freeman TC, Lenhard B, Bajic VB, Taylor MS, Makeev VJ, Sandelin A, Hume DA, Carninci P, Hayashizaki Y. (2014). A promoter-level mammalian expression atlas. Nature 507(7493), 462–470. doi: 10.1038/nature13182.
Haberle V, Li N, Hadzhiev Y, Plessy C, Previti C, Nepal C, Gehrig J, Dong X, Akalin A, Suzuki AM, van IJcken WFJ, Armant O, Ferg M, Strähle U, Carninci P, Müller F, Lenhard B. (2014). Two independent transcription initiation codes overlap on vertebrate core promoters. Nature 507(7492), 381–385.