Adria Canellas, who joined the Lymphocyte Development group at the LMS, has been awarded the Galen prize for the best MRes Biomedical Research student of 2016.
By Helen Figueira
December 22, 2016
Time to read: 4 minutes
Adria Canellas, who joined the Lymphocyte Development group at the LMS, has been awarded the Galen prize for the best MRes Biomedical Research student of 2016. His supervisor, Matthias Merkenschlager, said Adria achieved in a few months “the equivalent of a year’s work for most postdocs”. Here, Adria explains his work.
By Adria Canellas
The very fact that around 2 metres of DNA is packed into a nucleus with dimensions a million times smaller implies that despite the fact that genetic information is encoded in a linear sequence, the appropriate expression of genes must take place with the genome folded in three-dimensions. A key player in these interactions is cohesin, a ring-like complex that helps to orchestrate the folding of the genome and the expression of genes.
In 2013, the Lymphocyte Development Group had published that in cohesin-deficient cells, gene expression becomes more uniform. Highly expressed genes are preferentially downregulated, while those genes expressed at low level are upregulated. I wondered how this equalization of gene expression seen across populations would arise from individual cells. Perhaps the loss of cohesin might cause an increase of cell-to-cell variability in gene expression, for example because enhancer-promoter interactions might become more random.

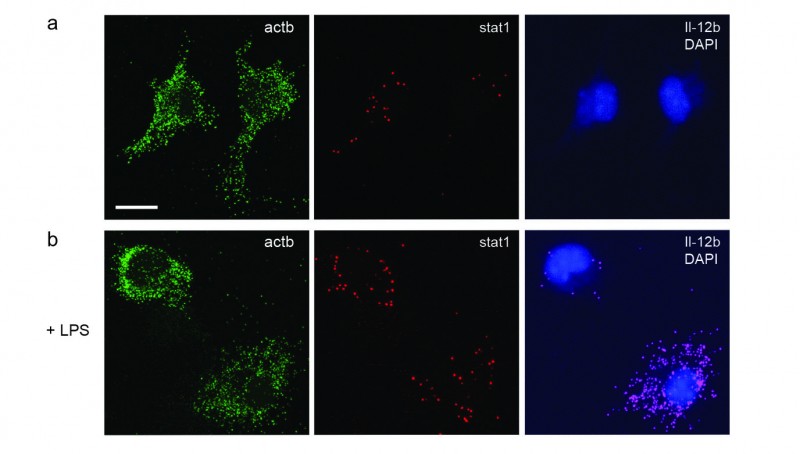
To answer this question, I implemented single-molecule RNA FISH (sm-FISH), a technique that quantifies gene expression in single cells with very little technical error. By imaging fluorescently labelled probes, sm-FISH allowed me simultaneously to visualize the mRNA molecules generated from up to three genes per experiment. When I combined this technique with a high-throughput image-analysis pipeline that enabled me to examine thousands of cells, I could measure the cell-to-cell variability in gene expression precisely.
Primary mouse macrophages provide a well-characterized model system for constitutive and inducible gene expression. I analyzed the expression of a housekeeping gene called Actb alongside the expression of Stat1 and Il12b, which are induced when macrophages are activated. My results suggested that cohesin deficiency did not strongly affect cell-to-cell variability in the expression of these genes in unstimulated cells. On the other hand, activated macrophages which lacked cohesin were unable to properly induce Il12b. Interestingly, this failure to upregulate Il12b was accompanied by an increase in cell-to-cell variability in Il12b expression.
Although this study needs to be extended to explore more genes before drawing definitive conclusions, my data provide preliminary evidence that cohesin could affect cell-to-cell variability or “noise” in gene expression. There is currently a great deal of interest in understanding the effect of this gene expression noise on the fitness of a multicellular organism. Many biological processes, such as the development of a zygote into a multicellular organism, require a high degree of coordination, and depend on tightly-regulated networks to read and express their genes at the appropriate moment and at the appropriate level. Could cohesin perhaps be playing the part of reducing gene expression noise in mammalian cells as they develop?
What I enjoyed the most about this project was working together with Dr. Matthias Merkenschlager and Dr. Sergi Cuartero, two brilliant, passionate and hard-working scientists. They were always available to help me, but also trusted in me and gave me freedom to explore, scientifically.
Building the high-throughput image-analysis pipeline was undoubtedly the most challenging part of my project. Even though I used recently published and freely available software developed by Professor Lucas Pelkman’s lab in the ETH Zurich, I had to spend many weeks figuring out how to make it work best in my system. Perhaps it was my perseverance and my fearlessness in this regard that earned me the Galen prize.
Postscript: Irene Robles-Rebollo, a PhD student with Merkenschlager (LMS) and Stumpf (Imperial College), has started combining single molecule RNA-FISH with mathematical modelling.