“What happens when meiosis goes wrong?”
Meiosis is the specialised cell division programme that produces reproductive cells known as gametes (sperm and eggs) that transmit the genome across generations. During meiosis, chromosome number must be halved to ensure that the correct number of chromosomes is restored upon fertilisation.
Our work focuses on conserved proteins that control the shape and function of meiotic chromosomes, including the cohesin complex and HORMA-domain proteins. We also investigate the quality-control mechanisms that monitor meiotic chromosome structure.
Defects that prevent the accurate partitioning of chromosomes during meiosis lead to infertility, birth defects and miscarriages in humans.
Our research aims to better understand the molecular mechanisms that ensure accurate chromosome halving during meiosis.
We use the nematode C. elegans and a combination of experimental approaches to investigate how proteins that control chromosome structure ensure accurate transmission of chromosomes during meiosis. We also exploit the genetic toolkit available in C. elegans to determine whether specific mutations identified in infertile patients cause infertility by hindering meiosis.
Our in vivo studies in C. elegans are complemented by biochemical and single-molecule imaging experiments using purified human proteins known to control meiotic chromosome structure.
Ultimately, our research aims to better understand the causes of human infertility and aneuploidy – cells with an abnormal number of chromosomes. Aneuploidy can lead to miscarriages and several to genetic disorders including Down syndrome.
We also aim to understand how alterations in the function of cohesin, a key determinant of chromosome structure, causes the developmental syndrome Cornelia de Lange.
“We investigate mechanisms of genome organisation and inheritance in germ cells to understand the causes of infertility and gamete aneuploidy, a leading cause of miscarriages and birth defects.”
Our group aims to understand the molecular mechanisms that ensure accurate chromosome segregation during meiosis, the specialised cell division programme that produces haploid gametes from diploid germ cells. Defects in this process cause infertility and are a leading cause of miscarriages and birth defects in humans. Using the nematode C. elegans as a model organism and a combination of experimental approaches, our research focuses on how different cohesin complexes and HORMA-domain proteins control the structure and function of meiotic chromosomes. We also investigate the quality-control mechanisms that orchestrate meiotic progression by monitoring meiotic chromosome structure. By focusing on these conserved components of the meiotic programme, our research aims to understand causes of human infertility and aneuploidy.
We are also investigating how cohesin complexes ensure normal organism development by controlling 3D genome organisation and gene expression, as mutations in cohesin are a cause of Cornelia de Lange syndrome.
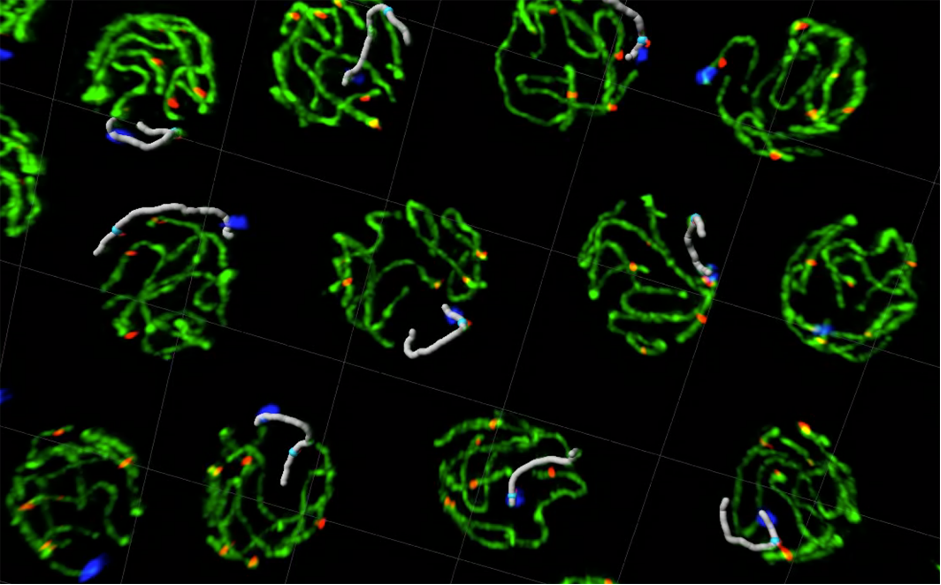
We investigate the mechanisms that ensure the accurate transmission of the genome from parents to offspring, which is essential for fertility and to prevent genetic diseases caused by aneuploidy such as Down’s syndrome.
We focus on understanding how proteins that bind DNA to control the structure and function of chromosomes ensure the successful completion of meiosis, the specialised cell-division program that produces haploid gametes from diploid germ cells. To this end, we combine multiple experimental approaches including biochemistry and single molecule imaging of meiotic proteins with in vivo studies of meiosis in the nematode C. elegans. We also use worms to determine if mutations identified in infertile patients impair meiosis.
In this way, we hope to understand the molecular mechanisms behind human infertility and to contribute to international efforts for the genetic diagnosis of infertility. This knowledge is required to design interventions aiming to restore fertility to patients carrying mutations that hinder the meiotic programme.
In vivo imaging of the meiotic divisions of a C. elegans oocyte in which chromosomes are labelled with a florescent protein. Note two consecutive rounds of chromosome segregation following by fusion with a sperm nucleus.

Castellano-Pozo M, Sioutas G, Barroso C, Lopez-Jimenez P, Jaso-Tamame AL, Crawley O, Shao N, Page J, Martinez-Perez E (2023). The kleisin subunit controls the function of meiotic cohesins by determining the mode of DNA binding and differential regulation by SCC-2 and WAPL-1. eLife 12:e84138 .
Láscarez-Lagunas LI, Nadarajan S, Martinez-Garcia M, Quinn JN, Todisco E, Thakkar T, Berson E, Eaford D, Crawley O, Montoya A, Faull P, Ferrandiz N, Barroso C, Labella S, Koury E, Smolikove S, Zetka M, Martinez-Perez E, Colaiácovo MP. (2022). ATM/ATR kinases link the synaptonemal complex and DNA double-strand break repair pathway choice. Curr Biol 32(21):4719-4726.e4.
Belan O, Barroso C, Kaczmarczyk A, Anand R, Federico S, O’Reilly N, Newton MD, Maeots E, Enchev RI, Martinez-Perez E, Rueda DS, Boulton SJ (2021). Single-molecule analysis reveals cooperative stimulation of Rad51 filament nucleation and growth by mediator proteins. Mol Cell 81(5):1058-1073.e7.
Castellano-Pozo M, Pacheco S, Sioutas G, Jaso-Tamame AL, Dore MH, Karimi MM, Martinez-Perez E (2020). Surveillance of cohesin-supported chromosome structure controls meiotic progression. Nat Commun 28;11(1):4345.
Link J, Paouneskou D, Velkova M, Daryabeigi A, Laos T, Labella S, Barroso C, Pacheco Pinol S, Montoya A, Kramer H, Markert S, Stigloher C, Martinez-Perez E, Dammermann A, Alsheimer M, Zetka M, and V. Jantsch (2018). Transient and partial nuclear lamina disruption promotes chromosome movement in early meiotic prophase. Dev Cell 45: 212-225.
Ferrandiz N, Barroso B, Telecan O, Shao N, Kim H-M, Testori S, Faull P, Cutillas P, Snijders AP, Colaiácovo MP and E. Martinez-Perez (2018) Spatiotemporal regulation of Aurora B recruitment ensures cohesion release during oocyte meiosis. Nat Commun 9: 838.