“Telomeres offer a unique ambivalence between protection of the chromosomes against genome instability and a challenging sequence for fork progression during DNA replication.”
DNA replication is an essential process for genome duplication, cell division and ultimately organismal survival that ensures faithful transmission of the genome to progeny. Certain genomic loci represent major obstacles to DNA replication including fragile sites, G-rich tracts and repetitive sequences, such as rDNA and telomeres. Mammalian telomeres have the propensity to adopt complex DNA secondary structures, including telomere-loops and telomeric G-quadruplex, which are believed to play essential roles in telomere maintenance. However, recent work has established that these structures are also a hindrance to DNA replication and might represent a potential source of genome instability, the hallmark of many diseases including cancer.
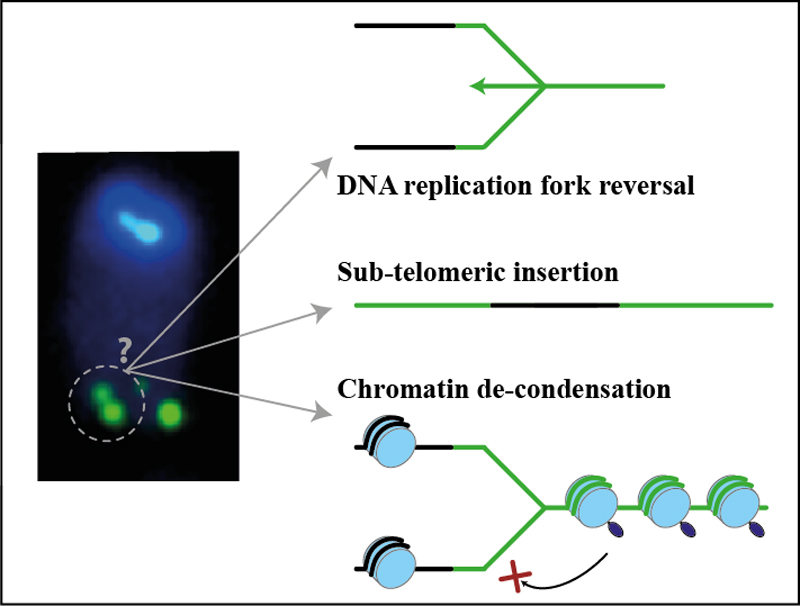
What is the nature of telomere fragility?
Despite recent advances, the mechanisms that facilitate DNA replication at mammalian telomeres remain unclear. It is important to address this question to decipher the mechanisms that help DNA replication through DNA secondary structures. Our group uses multidisciplinary approaches to investigate the cellular response to replication stress and the enzymatic activities that result in telomere replication aberrations, which involves direct visualisation of telomere abnormalities using complementary DNA-related methodologies and analysis of novel telomere-associated complexes. It is also critical to understand the structure of fragile telomeres, which remains poorly defined and represent a central question for the field using visualisation of biological molecules and proteomics.
The detailed investigation of the function of known and new factors that facilitate telomere DNA replication represent an outstanding challenge that will provide a novel framework for understanding the contributions of replication factors in general DNA replication, genome stability and cancer.
Selected Publications
SummersPA, Lewis BW, Gonzalez-Garcia J, Porreca RM, Lim AHM, Cadinu P, Martin-Pintado N, Mann DJ, Edel JB, Vannier J-B, Kuimova MK, Vilar R. (2021). Visualising G-quadruplex DNA dynamics in live cells by fluorescence lifetime imaging microscopy. Nature Communications, 12 (162).
Herrera-Moyano E, Porreca RM, Ranjha L, Gonzalez-Franco R, Skourti E, SunY, Stylianakis E, Montoya A, Kramer H and Vannier J-B. (2020). Human SKI is a telomere-associated complex involved in DNA-RNA hybrid control and telomere stability. BioRxiv: https://doi.org/10.1101/2020.05.20.107144
Summers PA, Lewis B, Gonzalez-Garcia J, Lim AHM, Cadinu P, Porreca RM, Martin-Pintado N, Mann D, Edel JB, Vannier JB, Kuimova MK and Vilar R. (2020) Visualising G-quadruplex DNA dynamics in live cells by fluorescence lifetime imaging microscopy. BioRxiv: https://doi.org/10.1101/2020.04.01.019794.
Porreca RM, Herrera-Moyano E, Skourti E, Law PP, Gonzalez-FrancoR, Montoya A, Faull P, Kramer H and Vannier J-B. (2020). TRF1 averts chromatin remodelling, recombination and replication dependent-Break Induced Replication at mouse telomeres. Elife 14, 9.
León-Ortiz AM, Panier S, Sarek G, Vannier JB, Patel H, Campbell PJ, Boulton SJ. (2018). A Distinct Class of Genome Rearrangements Driven by Illegitimate Recombination. Mol Cell.
Speckmann C, Sahoo SS, Rizzi M, Hirabayashi S, Karow A, Serwas NK, Hoemberg M, Damatova N, Schindler D, Vannier JB, Boulton SJ, Pannicke U, Göhring G, Thomay K, Verdu-Amoros JJ, Hauch H, Woessmann W, Escherich G, Laack E, Rindle L, Seidl M, Rensing-Ehl A, Lausch E, Jandrasits C, Strahm B, Schwarz K, Ehl SR, Niemeyer C, Boztug K, Wlodarski MW. (2017). Clinical and Molecular Heterogeneity of RTEL1 Deficiency. Front Immunol.
Sarek G, Vannier JB, Panier S, Petrini JH, Boulton SJ. (2015). TRF2 Recruits RTEL1 to Telomeres in S Phase to Promote T-Loop Unwinding. Mol Cell 57:4, 622–635 http://dx.doi.org/10.1016/j.molcel.2014.12.024.
Vannier JB, Sarek G, & Boulton SJ. (2014). RTEL1: functions of a disease-associated helicase. Trends in Cell Biology.
Vannier JB, Sandhu S, Petalcorin MI, Wu X, Nabi Z, Ding H, Boulton SJ. (2013). RTEL1 is a replisome-associated helicase that promotes telomere and genome-wide replication. Science 342(6155), 239–242.
Claudia HJ, Lin JR, Vannier JB, Slaats G, Kile AC, Paulsen, RD, Manning DK, Beier DR, Giles RH, Boulton SJ, Cimprich KA. (2013). NEK8 links the ATR-regulated replication stress response and s phase CDK activity to renal ciliopathies. Molecular Cell, 51(4), 423–439.
Ballew BJ, Joseph V, De S, Sarek G, Vannier JB, Stracker T, Schrader KA, Small TN, O’Reilly R, Manschreck C, Harlan Fleischut MM, Zhang L, Sullivan J, Stratton K, Yeager M, Jacobs K, Giri N, Alter BP, Boland J, Burdett L, Offit K, Boulton SJ, Savage SA, Petrini JH. (2013). A recessive founder mutation in regulator of telomere elongation helicase 1, RTEL1, underlies severe immunodeficiency and features of hoyeraal hreidarsson syndrome. PLoS Genetics 9(8).
Chapman JR, Barral P, Vannier JB, Borel V, Steger M, Tomas-Loba A, Sartori AA, Adams IR, Batista FD, Boulton SJ. (2013). RIF1 is essential for 53BP1-dependent nonhomologous end joining and suppression of DNA double-strand break resection. Molecular Cell 49(5), 858–871.
Vannier JB, Pavicic-Kaltenbrunner V, Petalcorin MI, Ding H, Boulton SJ. (2012). RTEL1 dismantles t loops and counteracts telomeric G4-DNA to maintain telomere integrity. Cell 149(4), 795–806.
Vannier JB, Depeiges A, White C, Gallego ME. (2009). ERCC1/XPF protects short telomeres from homologous recombination in arabidopsis thaliana. PLoS Genetics 5(2).
Vannier JB, Depeiges A, White C, Gallego ME. (2006). Two roles for rad50 in telomere maintenance. The EMBO Journal 25(19), 4577–4585.



