By Sophie Arthur
April 30, 2020
Time to read: 5 minutes
In 1858, the British anatomist Dr Henry Gray published the first pioneering and meticulous works describing how the human body is constructed. To this day, surgeons and clinicians use Gray’s blueprints to navigate the complex machinery of the human body, to know how it is connected, to diagnose and to repair our wounded or broken bodies. Now a century later, scientists are making blueprints that map out the structure of the hidden atomic world that underpins the functioning of every cell in our body, every bacterium and every virus.
This kind of scientific research, known as structural biology, uses sophisticated methods to determine the positions of every atom that make up the molecular machines that drive our cells. Where generally speaking, DNA is the genetic code that stores information for how a cell should work and proteins are the machines that are produced to carry out that work. These proteins include enzymes, cellular scaffolds, antibodies, the machinery for replicating DNA itself and the list goes on. These machines are structured in a way specialised to their task and the correct structure is essential to their proper function. Much like how the intricate anatomy of our organs are constructed to their individual tasks, so too the atoms of a protein are arranged specifically for that protein to work too.
Using structural biology and atomic descriptions of proteins, scientists can understand how these molecular machines are constructed and specialised to carry out their work. When a protein is mutated or damaged this often leads to a change in its structure and then an inability to carry out its function. Extremely detailed 3-dimensional atomic maps detailing these changes can help us understand how proteins misfunction in disease.
How is structural biology being used to fight COVID-19?
In the current COVID-19 pandemic, there is an incredible worldwide effort to tackle the disease. Part of this battle includes a worldwide community of structural biologists working together, sharing resources and at unprecedented speeds solving the structures of the proteins that make up SARS-CoV-2 virus. Cryo-electron microscopy (cryo-EM) is a structural biology technique that uses powerful electron microscopes to take hundreds of thousands of images of molecular machines frozen in ice at -180°C. Like a radiologist will make a 3D model of a patient from many 2D scans, the large numbers of images of a protein from a cryo-EM experiment are then put back together inside a computer to reconstruct the atomic structure of that protein. Using cryo-EM we now know the precise structure of the spike protein found on the outside of SARS-CoV-2 [1]. This protein allows the virus to enter and infect human cells.
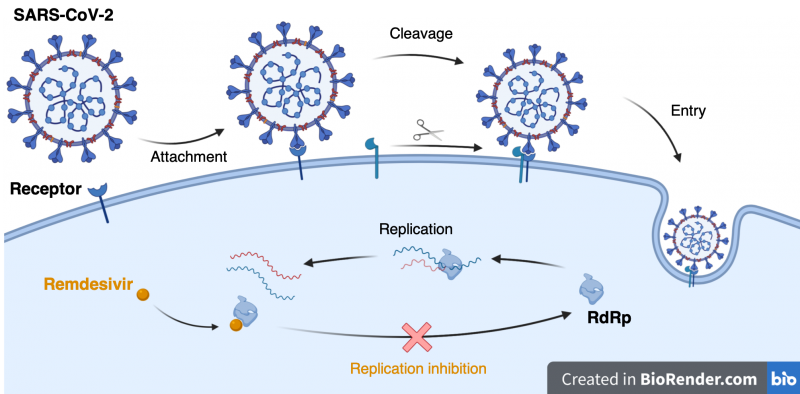
When the coronavirus spike protein engages with a human cell surface receptor, a cut is made by a human protein that opens up the viral spike protein. This causes a cascade of events that allow the virus to cross the outer membrane and into the cell. Scientists now know that this coronavirus has a change in its spike protein making it easier to cut, which might explain part of why SARS-CoV-2 is more infectious than previous coronavirus strains [2].
Knowing the structure of the spike protein provides the blueprint we need to target these mechanisms using drugs. Once inside the cell, the virus must replicate itself. The virus uses a protein called RNA dependent RNA polymerase (RdRp) to replicate its viral genome and is considered a major target for antiviral drugs such as Remdesivir. The atomic structure of coronavirus RdRp was also solved by cryo-EM [3] as part of the effort to understand the machines that replicate SARS-CoV-2 work. The structure of RdRp lets scientists see how drugs like Remdesivir target and stop coronavirus replication, leading the way to better drug design in the future. Taken together it is remarkable to witness the worldwide effort in response to the current pandemic. In particular, structural biology studies are revealing how coronavirus infection works and will provide the knowledge to build our molecular arsenal to fight this disease.
How is structural biology being used at the LMS?
At the MRC London Institute of Medical Sciences, researchers combine discovery and clinical science to understand the basis of healthy human development, cellular metabolism and function. Using structural biology, LMS scientists are unravelling the complex machinery underlying cellular events that are fundamental to human life and of high therapeutic importance. Proteins that replicate human DNA, the machinery that maintains genome stability, chromatin structure, chromosome integrity and future technologies in genome editing are all but a few of the targets we aim to understand structurally. By understanding the proteins that govern genetic programs, control chromatin structure and enact epigenetic memory we are pushing discovery science which combined with medical insight will drive innovation and health.
References:
Article and images by Kyle Morris, Senior Investigator Scientist, MRC LMS. @kylelmorris
Title image credit: Gray’s anatomy 1918, NIAID-RML, Walls et al 2020 Cell